Enzyme cost minimization
Main | Workflow | Model | Running ECM | Download
|
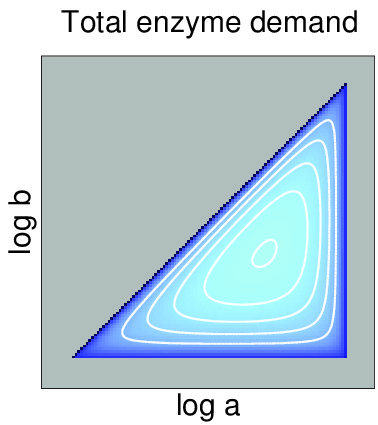
Enzyme cost minimization Enzyme cost plays a major role in the choice of
metabolic routes, both in evolution and
bioengineering. If the desired fluxes are known, minimal
requires enzyme levels, as well as corresponding
metabolite levels, can be predicted based on known rate
laws and on a principle of minimal enzyme cost. |
Aside from being a practical simulation method, ECM also
yields some theoretical insights. The optimal enzyme profiles depend on kinetic rates laws and
are therefore shaped by factors such as catalytic
constants, thermodynamic driving forces, enzyme
saturation, and allosteric regulation. The enzymatic cost
can be split into multiplicative contributions from enzyme
capacity, energetics, and kinetics. By neglecting some of
these factors, we construct simplified enzyme cost
functions, applicable even if few or no parameter values
are known. The energetic cost contribution is directly
linked to thermodynamic constraints: if a reaction is
thermodynamically unfeasible, we can simply see this as a
case of infinitely high enzyme cost.
The enzyme costs computed by ECM lead to general flux cost
functions, which can be used to bridge the gap between
constraint-based and kinetic metabolic models. Finally, the
optimality condition of ECM can be interpreted as a balance
relation between the costs of adjacent enzymes. This links flux
analysis to metabolic control analysis.
On this website, you can find software for enzyme cost minimization (in matlab and python), as well as data files for an illustrative example, a kinetic model of central carbon metabolism in E. coli.
References
If you use enzyme cost minimization in your work, please cite our articles
Noor E., Flamholz A., Bar-Even A., Davidi D., Milo R.,
Liebermeister W. (2016), The protein cost of metabolic fluxes:
prediction from enzymatic rate laws and cost minimization,
PLoS Comp. Biol. 12 (10): e1005167
Metabolic enzyme cost explains variable trade-offs
between microbial growth rate and yield, Wortel M.T., Noor
E., Ferris M., Bruggeman F.J., Liebermeister W. (2018), PLoS
Computational Biology 14(2): e1006010
Contact
Please contact Wolfram Liebermeister or Elad Noor with any questions or comments.